tNodeEmbed — Node Embedding over Temporal Graphs
By Uriel Singer
In this post I’ll take about: tNodeEmbed (joint work with Dr. Ido Guy & Prof. Kira Radinsky). tNodeEmbed is a method that leverages the temporal information in graphs in-order to create richer node embedding representations. This concept was written up as a paper which was accepted to IJCAI2019.
Let’s start talking about embeddings in the graph domain.
Embeddings in the graph domain
They are many ways to do embeddings in the graph domain:
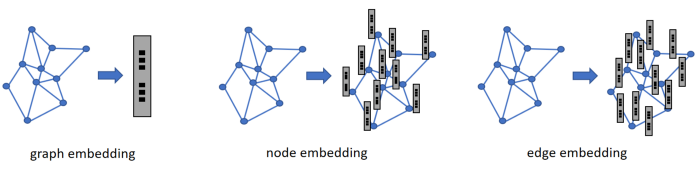
In Figure 1 we can see that it is possible to embed the whole graph, embed each node and embed each edge (one naive way is by merging it’s two node embeddings).
All of these embeddings can be then used in-order to answer different ML questions.
Temporal graphs hold rich information that can’t be captured from a static graph. For example, suppose Eve became Facebook friends with Alice and Bob in the same day, while she became friends with Chris a year ago. Who is more likely to become friends with Alice? is it Bob or Chris? In a static graph there is no difference (4 nodes with 3 edges: Eve to Alice, Bob and Chris). A temporal graph holds the information on when each edge occurred, which resolves us with the conclusion that Alice is more likely to become friends with Bob than Chris.
The Idea behind this paper is learning Node Embedding over Temporal Graphs:
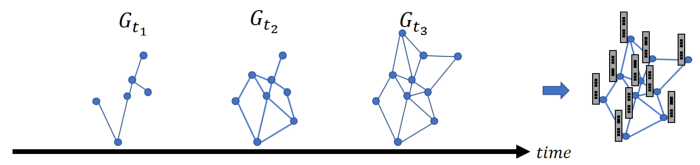
Figure 2 illustrates the idea behind node embedding over temporal graphs. We define the graph in time t to be the graph that holds all edges which appeared at any point up to time t.
tNodeEmbed
Lets start by overviewing tNodeEmbed:

tNodeEmbed consists from 3 steps:
1. Initialization
Several prior works studied node embedding and reported good performance on static graphs. The architecture presented in this paper can use any of the known approaches for node embedding over static graphs. Specifically, we opted to work with node2vec, which achieved state-of-the-art performance on various benchmarks.
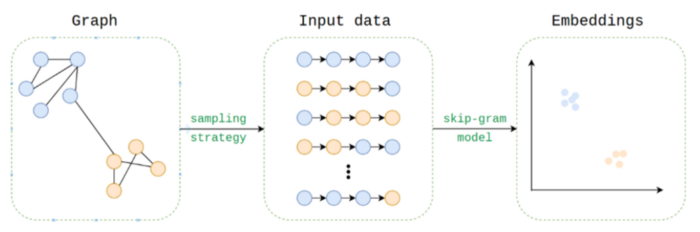
In Figure 3 we can see the node2vec overview, which is simply a reduction from the graph domain to the text domain: Each node is treated as word while the ‘sentences’ are generated by sampling many random walks. Those ‘sentences’ are inserted into a word2vec model. The word embeddings outputs are returned to the graph domain as the nodes embeddings.
We use node2vec to initialize Qt. Specifically, we compute the node embeddings for all T graphs Gt1, …, GtT. The outcome of this stage is a d X Tvector per node, where T is the number of time steps and d is the embedding size (each column is a different static embedding).
2. Temporal Node Embedding Alignment
In-order for us to be able to look on how a specific node moved between the time-steps in the embedding space, we still need to align the time-steps embeddings.
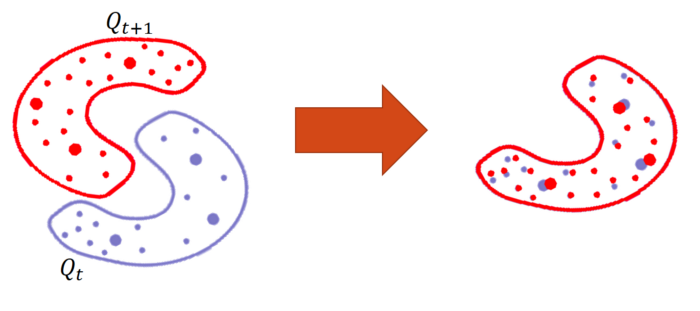
In order to do so, we apply Orthogonal Procrustes on every two consecutive time-steps iteratively. The assumption we use here is that the 2 embedding spaces are close to each other enough for them to be aligned by a rotation matrix. This is a reasonable assumption due to that we do it on 2 consecutive time-steps that differ only in the edges that were addedbetween the two time-steps.
For people that find this technique interesting, here’s another research by FAIR which uses the same approach for unsupervised NMT.
3. Node Embedding over Time
In the final step we merge the d X T vector of each node to a d vector using a LSTM. The LSTM is then connected into an additional layer that tries to predict a specific task is can be seen in Figure 6.

Results
We compared tNodeEmbed to various baselines, both static and temporal, and on a various different datasets. Two tasks we experimented on are the temporal link-prediction and multi-label node classification tasks:
1. Temporal Link Prediction
In-order to create this task, we picked a pivot-time step in such a way that all edges that are known up to that pivot time-step are used for training while the edges that are known after that pivot time-step are used for test.

An interesting observation is that the lower the clustering coefficient (CC), the higher the gap between tNodeEmbed to its static baseline, node2vec.
We understand from this phenomena that the “denser” the graph, the less informative the temporal information is in-order to predict future links. The reason lies in the fact that edges tend to attach to higher degree nodes, and in “denser” graphs, these nodes have extremely high degrees compared to other nodes.
2. Multi-Label Node Classification

In Figure 2 we can observe the tNodeEmbed does pretty well.
Final Words
tNodeEmbed nailed it!

Thank you for reading. Feel free to use this work or continue the research and contribute to the graph embedding community!
Important Links
The paper: https://www.ijcai.org/proceedings/2019/0640.pdf
The code: https://github.com/urielsinger/tNodeEmbed


Why visitors still use to read news papers when in this technological world the whole
thing is available on web?
I could not resist commenting. Well written!
If you are going for finest contents like myself,
simply pay a quick visit this site every day because it presents feature contents, thanks